More well thought out work can be found at — https://axial.substack.com/
Axial partners with great founders and inventors. We invest in early-stage life sciences companies often when they are no more than an idea. We are fanatical about helping the rare inventor who is compelled to build their own enduring business. If you or someone you know has a great idea or company in life sciences, Axial would be excited to get to know you and possibly invest in your vision and company . We are excited to be in business with you - email us at info@axialvc.com
Inventors #15
A set of ideas and observations on inventions and discoveries in life sciences.
Immunology
The immune system and everything in it.
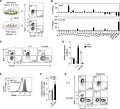
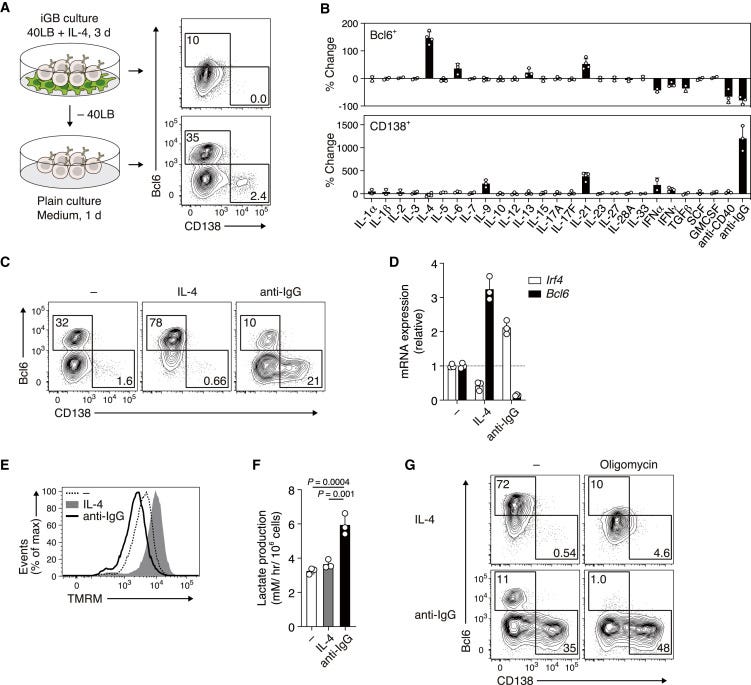
Metabolic Reprogramming Induces Germinal Center B Cell Differentiation through Bcl6 Locus Remodeling - https://www.cell.com/cell-reports/fulltext/S2211-1247(20)31322-X - the Kitamura Lab at Tokyo University of Science discovered the IL-4 is important to reprogram the metabolism of B-cells to promote their differentiation in germinal centers (GC):
IL-4 is a cytokine known to activate B-cell and promote differentiation to plasma cells (shift to antibody secretion)
GCs are sites found within lymph nodes and the spleen where activated B-cells undergo somatic hypermutation (SHM) to generate high-affinity antibodies
Bcl6 is the master regulator of B-cells becoming activated and migrating to a GC
The group sought to understand the role of B-cell metabolism during this process of activation and SHM
Using their iGB culture method, the group screened for cytokines that convert Bcl6− cells into Bcl6+ cells. Expectedly, they found that IL-4 was the most potent cytokine among IL-6/13/21 and was found to increase mitochondrial membrane potential.
With the latter discovery, the metabolome was profiled in Bcl6+ cells after IL-4 stimulation - αKG, a metabolite important for the citric acid cycle (TCA) and a cofactor for H3K27-demethylase, an enzyme to remove the repressive epigenetic marker. They found that IL-4 reduced H3K27m3 markers in the Bcl6 locus
Ultimately, the group used a series of simple and elegant experiments to connect metabolic reprogramming to Bcl6 and B-cell differentiation. B-cell metabolism could become a new target for a wide range of immunological disorders.
Biochemistry and structural biology
The granddaddy of them all.
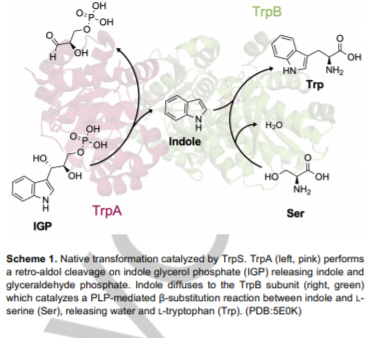
Tryptophan Synthase: Biocatalyst Extraordinaire - https://chemistry-europe.onlinelibrary.wiley.com/doi/10.1002/cbic.202000379 - a useful review out of the Arnold Lab at Caltech on tryptophan synthases (TrpS):
Tryptophan synthases (TrpS) is an enzyme involved in the biosynthesis of tryptophan
The enzyme catalyzes carbon-carbon (C-C) bonds between an indole and serine to make Trp. With this unique capability, TrpS has become a workhorse to produce noncanonical amino acids (ncAA) and as a biocatalyst for new reactions.
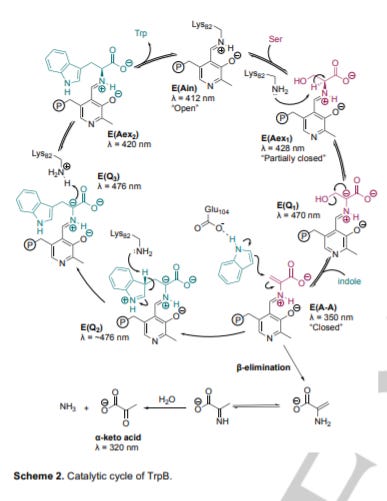
This is enabled by TrpS’ flexibility for indole analogs; however, more work is needed to unlock the enzymes constraints to expand its potential substrates. A key breakthrough was the use of directed evolution to develop a standalone TrpS β-subunit (TrpB) and free it from the allosteric regulation by the alpha subunit of TrpS. This immediately opened up the number of substrates TrpS can recognize.
The two major applications of TrpS design are: ncAA synthesis and biocatalytic cascades
ncAA synthesis - creating noncanonical amino acids and expanding the number of indole analogs as a substrate
Biocatalytic cascades - synthesizing D-amino acids and Trp derivatives- https://pubs.acs.org/doi/10.1021/acscatal.9b00739 and https://chemistry-europe.onlinelibrary.wiley.com/doi/abs/10.1002/cbic.201900069
TrpS is a scaffold for enzyme design, and the recent use of TrpB variants has the potential to create new chemistry and transformative products
Neuroscience
Roughly 20 years behind but set up to transform the concept of human.
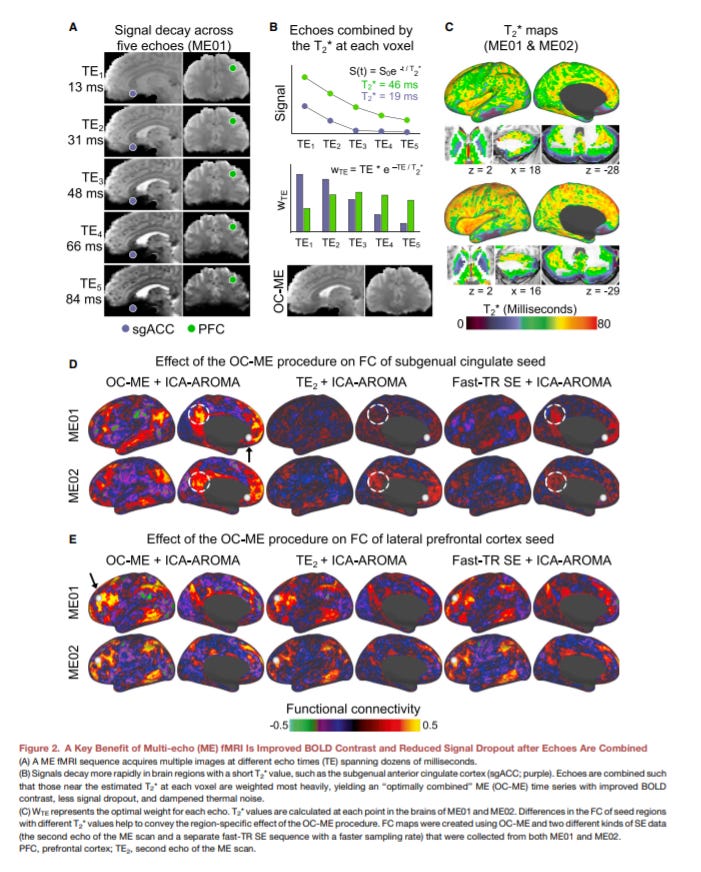
Rapid Precision Functional Mapping of Individuals Using Multi-Echo fMRI - https://www.cell.com/cell-reports/fulltext/S2211-1247(20)31529-1 - the Liston Lab at Cornell showed that multi-echo fMRI creates more accurate brain connectivity data for individual patients:
fMRI is mostly done with a single-echo method where a single brain image is collected after a radio frequency (RF) pulse. Mult-echo collects multiple images at varying volumes per RF - https://fim.nimh.nih.gov/projects/multi-echo-fmri
Multi-echo fMRI improves the fidelity of brain connectivity maps with the tradeoff on full-brain coverage. New technologies are making this compromise less acute: (1) single-echo has full-brain coverage or complete spatiotemporal resolution and has longer time scans (i.e. ~30 minutes) and (2) multi-echo is more sensitive and has shorter time scans (i.e. ~10 minutes)
The paper shows that the method has higher sensitivity for specific regions of the human brain: subgenual cingulate, basal ganglia, and cerebellum
With this discovery, the group showed that multi-echo fMRI has the potential to reliably compare individual brain connectivity maps by focusing on differences within the 3 regions. The potential is to use the tool for neuroimaging across large populations to discover new networks that drive everything from awareness to neurological disorders.
The key breakthroughs are: (1) finding individual differences in fMRI scans with less data and (2) develop a clinically useful method for neuroimaging
Cell biology
Cell structure and function.
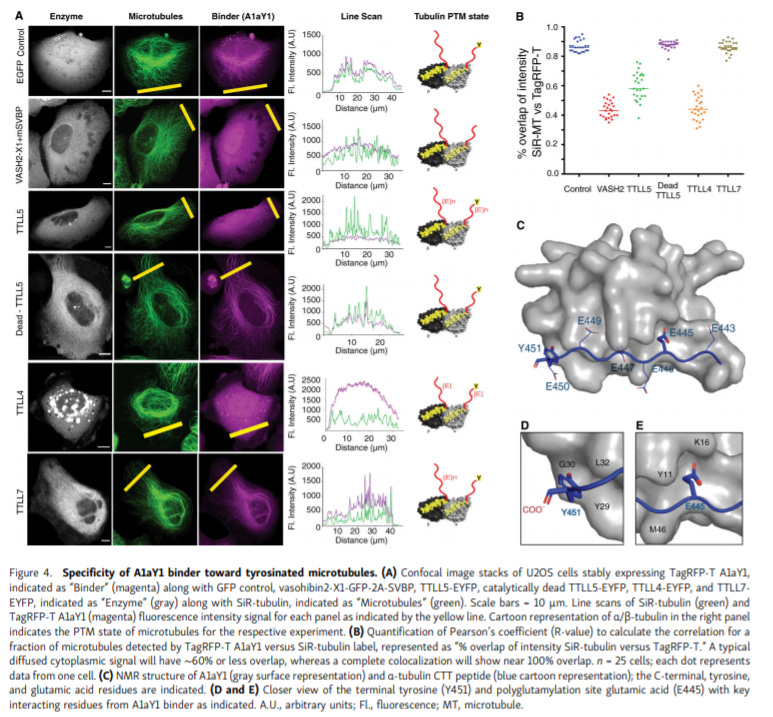
Genetically encoded live-cell sensor for tyrosinated microtubules - https://rupress.org/jcb/article/219/10/e201912107/152071/Genetically-encoded-live-cell-sensor-for - the Sirajuddin Lab at inStem invented a new genetically encoded nanobody-based sensor to detect tyrosinated microtubules in living cells:
Microtubules determine cell dynamics and organization. Developing new tools to measure tubulin post-translational modifications (PTM) has the potential to unveil new areas of cell biology and identify new pathways important for microtubule stability
The group used a yeast display method to identify a hit for terminal tyrosine on α-tubulin. After optimization, a single-domain antibody (i.e. nanobody) was developed as a specific sensors for tyrosinated microtubules in live cells
The group then chemically induced microtubule depolymerization to validate that their sensor worked
Like all sensors, this invention opens up avenues to study the effects of PTMs on microtubules and discover new drug candidates centered around cell dynamics
Next steps are to bring the same method to identify sensors for other microtubule PTMs - acetylation, glutamylation, and glycylation
Genetics, genomics, and developmental biology
Heredity and variation.
Perivascular mesenchymal cells control adipose-tissue macrophage accrual in obesity - https://www.nature.com/articles/s42255-020-00301-7 - out of the Gupta Lab at University of Texas Southwestern Medical Center, the group connects a “new” cell population in white adipose tissue (WAT) to inflammation and macrophage accumulation:
Inflammation in WAT is a symptom of obesity. Fibro-inflammatory progenitors (FIP) were recently discovered to activate inflammatory pathways are high-fat diets in mice - https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6167054/ This paper is a next step to understand this mechanism.
The group profiled the gene expression of FIPs under a high-fat diet regimen. They found that activation of the cell type was driven by the downregulation of a NF-κB signaling repression - zinc-finger protein 423 (ZFP423)
To verify this, ZFP423 expression induced in activated FIPs reduced WAT inflammation and other proinflammatory markers
Using mouse model, the paper maps out the mechanism of FIPs and identifies ZFP423 as a central mediator of the transition to a pro-inflammatory environment in WAT
Next steps are to understand the triggers for this response. Gut-derived antigens? Lipids from food? What other cell types coordinate with FIPs?