More well thought out work can be found at — https://axial.substack.com/
Axial is building a new company in kidney disease. If you want to get involved to transform the lives of patients, apply here: https://biofounder.axialvc.com/ We are looking for potential founders, builders, and inventors with interests to combine biology with software. Our goals are to help the millions of kidney disease patients and build an enduring business.
Inventors #23
A set of ideas and observations on inventions and discoveries in life sciences.
Immunology
The immune system and everything in it.
Central memory CD8+ T cells derive from stem-like Tcf7hi effector cells in the absence of cytotoxic differentiation - https://www.cell.com/immunity/fulltext/S1074-7613(20)30399-X - the Held Lab at UNIL discovered a new population of CD8+ (killer) T-cells that retained many stem-line features even after differentiation:
Transcription factor T-cell factor 1 (TCF1) is encoded by the Tcf7 gene and is regulated by the Wnt pathway. The group’s premise was to study the role of TCF1 in response to acute viral infections.
A Tcf7GFP reporter was engineered into CD8+ T-cells and transplanted into a mouse model. One day post-transplant, mice were infected with lymphocytic choriomeningitis virus (LCMV).
They discovered the Tcf7 expression was able to sort memory T-cells into 3 classes: Tcf7GFPHi CD62L-positive (another marker for memory cells), Tcf7GFPHi CD62L-, and Tcf7GFPLo CD62L-.
From these 3 classes, Tcf7 expression was indicative of a T-cell’s ability to hone into lymph nodes and expanded 1000-fold within a week of the infection. Whereas, Tcf7GFPlo cells mimicked the expression profiles based on increased expression of GzmA, CX3CR1 and decreased for CD127 of effector memory T-cells (Tem) that are short lived.
The paper established the killing activity of a new T-cell subset and validated a new biomarker, in mice, to differentiated memory T-cells.
Biochemistry and structural biology
The granddaddy of them all.
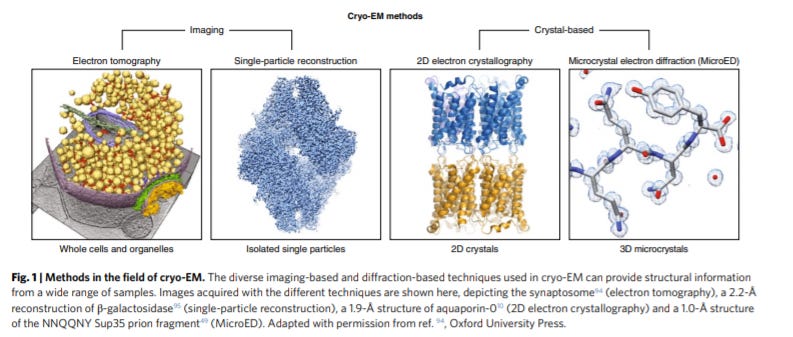
The cryo-EM method microcrystal electron diffraction (MicroED) - https://www.nature.com/articles/s41592-019-0395-x - a review led by the Gonen Lab at UCLA does a great job at giving an overview on microcrystal electron diffraction (MicroED) to determine protein structures:
The group invented MicroED in 2013 to generate diffraction data from crystals with ~5x lower volumes. The method relies on standard equipment but relies on collecting data from low electron doses and indexing. So the ability to determine structures from microcrystals was a breakthrough for large complexes and membrane proteins.
Since 2013, data analysis has become a lot more accessible and cryo-EM devices more precise creating a golden age for structural biology
2D electron crystallography for cryo-Em is an important approach where the advantages are that useful data can be generated from a single diffraction pattern, but there are limits on sample input size. MicroED focused on 3D methods that are more data-intensive and require several patterns, but this complexity enables smaller inputs. MicroED was the first to show the ability to get structural from microcrystals with a PoC for the lysozyme structure to 2.9-Å resolution.
In 2014, the group added continuous rotation to MicroED to speed up this data collection process and increase sampling rates. With this improvement, MicroED was able to determine the lysozyme structure at a 2.5 Å resolution. Better data processing got this to a 1.8-Å resolution.
The real unique advantage of MicroED is its processing speed and ability to study protein dynamics across time, not just structures. Things like enzyme reactions and ligand binding are perfect fits for this tool.
Cryo-EM applications are still growing. Structures are being determined at a faster rate, but the larger opportunity is then translating these tools to drug development.
Neuroscience
Roughly 20 years behind but set up to transform the concept of human.
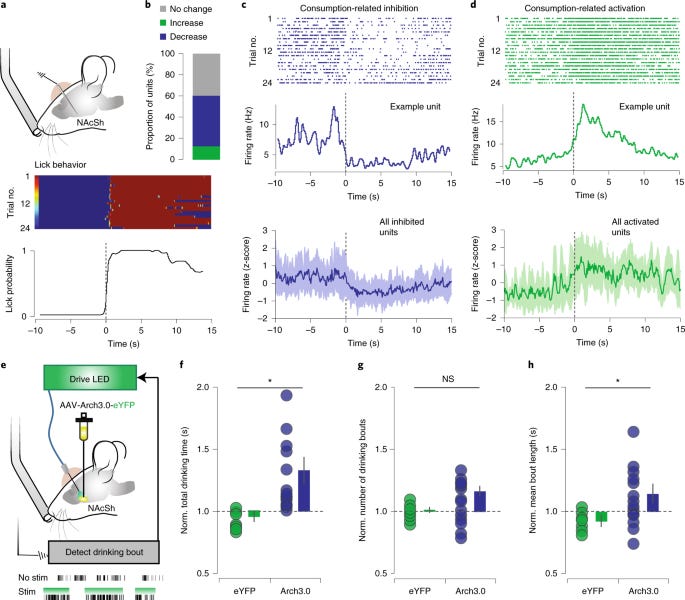
Ventral arkypallidal neurons modulate accumbal firing to promote reward consumption - https://www.nature.com/articles/s41593-020-00772-7 - out of the Creed Lab at WUSTL, the group discovered a new population of ventral pallidal (VP; a key part of the reward system in the brain) neurons that modulate neurons that were thought to be upstream of VP:
VP neurons are important for creating reward values in the brain. The nucleus accumbens shell (NAcSh) class of neurons that are known to be inhibited during reward consumption.
The paper used a mouse model to measure reward consumption based on drinking from a lickometer. During this process, over a third of NAcSh neurons were inhibited. Optogenetic inhibition of these neurons was also sufficient to induce a reward response.
The group hypothesized that VP neurons could be acting on NAcSh neurons. It’s not clear how they got to this idea; maybe screening a wide set of neurons?
To confirm VPs roles in this process, AAV-Cre constructs were used to label NAcSh neurons and a Cre-dependent eYFP reporter construct into VP neurons. This experiment confirmed VPs projection into NAcSHs; the group named these neurons as part of the ventral arkypallidal (vArky) class.
The work discovered a unique subpopulation of neurons that act in a unique way to sustain reward consumption.
Viral tracing has been such a powerful tool to study the CNS, and discoveries like these are important to understand what drives reward-seeking behavior.
Cell biology
Cell structure and function.
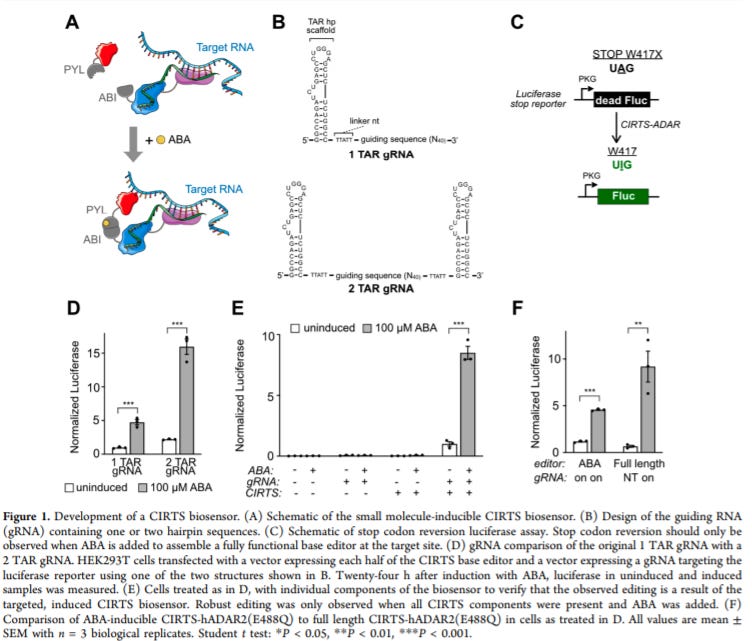
Small Molecule-Inducible RNA-Targeting Systems for Temporal Control of RNA Regulation - https://pubs.acs.org/doi/10.1021/acscentsci.0c00537 - the Dickinson Lab at the University of Chicago invented a small molecule-inducible CRISPR/Cas-inspired RNA targeting system (CIRTS):
Previously, the group had invented CIRTS to use CRISPR effectors to target RNA for a wide range of modifications as well as selective degradation and translation.
This paper developed effectors that are inducible by small molecules to imbue both spatial and temporal control of the tool.
To add the small molecule inducing feature, the group combined CIRTS with heterodimerization domains (ABI and PYL) of the abscisic acid (ABA), which is mostly orthogonal to mammalian targets. ABI was fused to the RNA binding portion of the CRISPR effector and PYL was fused to the effector domain. ABA would then enable complex formation and activity.
As a PoC, the group tested their ABA-CIRTS system on a luciferase mRNA with a premature stop codon that can be turned back on through A-to-I editing. They also validated the tool in vivo.
RNA editing either through CRISPR or ADARs is a useful tool and has the potential to become medicines down the line.
Genetics, genomics, and developmental biology
Heredity and variation.
Design of proteins presenting discontinuous functional sites using deep learning - https://www.biorxiv.org/content/10.1101/2020.11.29.402743v1 - out of the Baker Lab at the University of Washington, the group used a method called deep network hallucination to design new proteins without any prior constraints:
The key premise is to “hallucinate” new proteins that do not have backbone constraints. This starting point allows machine learning models to get to a successful endpoint - a binder for a target.
This application of a sampling method to protein design enables the ability to screen different protein structures for very discontinuous target surfaces. Rather than pre-specifying a backbone, this method allows a model to find the optimal backbone.
The group chose complement cascade protein C3d, IL-10, and IL-4 as PoCs to generate mimetic with similar functions that were validated in silico.
The key value here is that the method doesn’t need to know arrangements of functional sites beforehand to design de novo protein candidates.